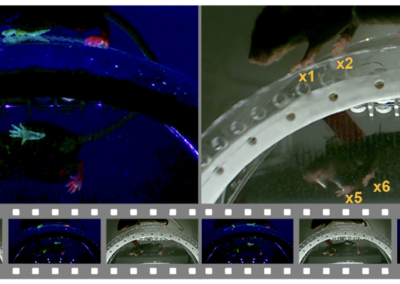
Several popular methods for pose estimation have emerged in recent years. DeepLabCut and LEAP are popular tools for tracking actions of individual animals. The projects are well documented and numerous tutorials exist that make it easy for new users to learn and use the software. The packages were developed by neuroscience research groups. The papers reporting the methods received a lot of attention on social media and have been widely cited. An alternative method is DeepPoseKit. It was developed by behavioral scientists at the Max Planck Institute of Animal Behavior. The lead developer is Dr. Jacob Graving. The research team published a paper in eLife in 2019 reporting several innovations that reduce training time while retaining high accuracy. A major innovation of their approach was the development of the Stacked DenseNet for pose estimation. The paper is very clearly written, and provides a comprehensive introduction to the main computational concepts that underlie pose estimation and tracking. The published appendixes are pure gold for anyone interested in learning about these methods. In our experiences in using the package, it is easy to use. There are clearly written tutorials. The code is standard for the Python universe, and the functions have easily accessible key word arguments that make it easy to change parameters. Outputs are saved in easily accessed files. The package is able to use labeling from the excellent DeepLabCut GUI, so you don’t need to start from scratch, and can replicate the analysis approach used by both DeepLabCut and LEAP. If you are just getting started with pose estimation and tracking or wish to try out an alternative approach, DeepPoseKit is very much worth your time. It is also an excellent tool for teaching current methods for video analysis to trainees, especially given the very clearly written paper on the toolkit and very accessibly API. This research tool was created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the tool was described, and include an RRID in the Materials and Methods of your future publications. Project portal RRID:SCR_021395; Software RRID:SCR_021405 Get access to DeepPoseKit files and instructions on Github. Read more about DeepPoseKit from the recent eLife publication!DeepPoseKit
GitHub Repository
Read the Paper
Have questions? Send us an email!






