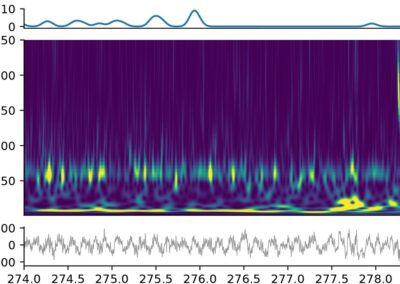
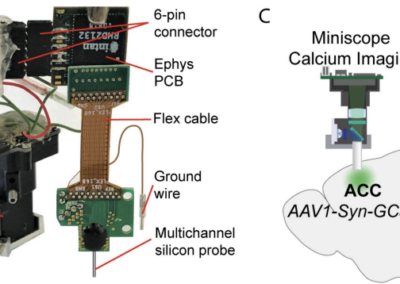
Peter Petersen and team have published an article to Neuron detailing their innovative framework CellExplorer. This open-source technology aims to facilitate and standardize the classification of single neurons throughout various laboratories. Neuronal cell type classification varies across labs, which can limit the ability of researchers to compare single unit electrophyisiological data from multiple research groups. In order to identify possible correlates between various neurons and their functions, a standardized framework that allows for the classification and logging of findings is required, which is what CellExplorer hopes to achieve. An open-source tool written primarily for C and Python, CellExplorer’s MATLAB based framework is based on three components: a processing module, a flexible data structure, and a graphical interface component. The first component aims to provide a standardized data calculation based on physiological metrics. The second component allows for easy storage of processed data. The final component provides a visual representation of the standardized data calculations and can compare these findings with findings reported in other labs to encourage the comparison of the metrics. The article includes examples of the visual representations of data that the CellExplorer program was able to identify across three different labs. This research tool was created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the tool was described, and include an RRID in the Materials and Methods of your future publications. Project portal RRID:SCR_022358 The full paper including extensive examples of the data the program has been able to identify can be found here! Get access to the software, docs, and tutorials from the CellExplorer Docs page! This post was brought to you by Zoe Joy. This project summary is a part of the collection from neuroscience undergraduate and graduate students in the Computational Methods course at American University.CellExplorer

Read the Paper
CellExplorer Documentation
Thanks, Zoe!
Have questions? Send us an email!